28 May 2010
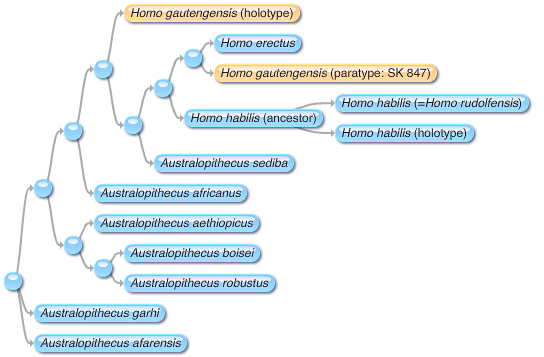
gautengensis in the sediba phylogeny
A Homo gautengensis by any other name...
Curnoe (2010). A review of early Homo in southern Africa focusing on cranial, mandibular and dental remains, with the description of a new species (Homo gautengensis sp. nov.). HOMO Journal of Comparative Human Biology (online early). doi:10.1016/j.jchb.2010.04.002
Here's the abstract:
The southern African sample of early Homo is playing an increasingly important role in understanding the origins, diversity and adaptations of the human genus. Yet, the affinities and classification of these remains continue to be in a state of flux. The southern African sample derives from five karstic palaeocave localities and represents more than one-third of the total African sample for this group; sampling an even wider range of anatomical regions than the eastern African collection. Morphological and phenetic comparisons of southern African specimens covering dental, mandibular and cranial remains demonstrate this sample to contain a species distinct from known early Homo taxa. The new species Homo gautengensis sp. nov. is described herein: type specimen Stw 53; Paratypes SE 255, SE 1508, Stw 19b/33, Stw 75–79, Stw 80, Stw 84, Stw 151, SK 15, SK 27, SK 45, SK 847, SKX 257/258, SKX 267/268, SKX 339, SKX 610, SKW 3114 and DNH 70. H. gautengensis is identified from fossils recovered at three palaeocave localities with current best ages spanning ~2.0 to 1.26–0.82 million years BP. Thus, H. gautengensisis probably the earliest recognised species in the human genus and its longevity is apparently well in excess of H. habilis.The holotype, Stw 53, has previously been referred to either Homo habilis Leakey & al. 1964 or Australopithecus africanus Dart 1923. Interestingly, though, one of the paratypes, SK 15, is already the holotype of Telanthropus capensis Robinson 1949! So Homo gautengensis would appear to be a junior subjective synonym.
It gets even more interesting (for us nomenclature buffs, anyway): if Telanthropus capensis were to be transferred to Homo, then it would be a junior homonym of Homo capensis Broom 1918 (a.k.a. "Boskop Man"), which itself is a junior subjective synonym of Homo sapiens (the type specimen probably representing an early Khoisan individual).
I'm not sure if any of this is discussed by Curnoe, because I don't have access to the paper. If anyone has a PDF, feel free to e-mail to keesey [at] gmail [dot] com.
UPDATE: I have the paper now and will be looking it over.
ANOTHER UPDATE: Telanthropus is mentioned in passing, but the synonymy is not discussed.
27 May 2010
Upcoming Names on Nodes Presentation
I have my work cut out for me....Names on Nodes: Automating the Application of Taxonomic Names within a Phylogenetic Context
Names on Nodes1 is an open-source2 Flex application which utilizes a mathematical approach to automate the application of phylogenetically-defined names to phylogenetic hypotheses. Phylogenetic hypotheses are modeled as directed, acyclic graphs, and may be read from bioinformatics or graph files (Nexus, NexML, Newick, and GraphML) or created de novo. Hypotheses may also be merged from multiple sources. Names on Nodes stores hypotheses as MathML, an XML-based language for representing mathematical content and presentation. Phylogenetic definitions may be constructed using a visual editor and exported in MathML. Thus, it is possible to create a dictionary of defined names and automatically apply them to phylogenetic hypotheses. In the current version of the application, such dictionaries exist only as MathML files, but in future versions definitions may also be loaded from databases (e.g., RegNum).
Additional functionality in Names on Nodes includes the ability to coarsen a phylogenetic graph (thereby simplifying it while still reflecting the overall structure) or to export it as an image file (raster or vector, potentially with semantic annotations).
- Source code available at: http://bitbucket.org/keesey/namesonnodes-sa/
- MIT license
26 May 2010
Upcoming Talk: Toward a Complete Phyloreferencing Language
Toward a Complete Phyloreferencing Language
A phyloreference is a statement indicating a taxon within a phylogenetic context. A common use for phyloreferences is in phylogenetic definitions, which tie taxonomic names to taxa via such statements. Several conventions for writing phyloreferences have been proposed, but most only cover a few “standard” forms (node-, branch-, and perhaps apomorphy-based clades) without the capacity to represent more “exotic” forms (e.g., ancestor-based clades and qualified/modified references). In order to build a complete phyloreferencing language, the mathematical underpinnings of phylogenetic contexts must be clarified. A phylogenetic context may be modeled as a directed, acyclic graph, wherein nodes model taxonomic units and directed edges model immediate descent. Higher taxa are modeled as unions of nodes. A phyloreferencing language must minimally allow for certain classes of entity: Boolean values, sets (including taxa, relations, and the empty set), and lists (including graphs and functions). It must also minimally allow for basic operations related to logic, set theory, and graph theory. Higher structures such as declarations and piecewise constructs must also be possible. With these as a basis, functions related to phylogeny can be defined: maximal, minimal, predecessor union/intersection, successor union/intersection, exclusive predecessors, synapomorphic predecessors, clade, crown clade, and total clade. I show how such a language may be used to represent various types of phyloreference, both “standard” and “exotic”.
Now to figure out how to condense that into a five-minute talk....
21 May 2010
Names on Nodes Issue Tracker
Essential features left to implement, complete or fix:
FILES AND FORMATS
- Certain formats for import, especially NexML and NEXUS. (Currently only Newick can be imported. MathML files can be loaded as well.)
- Certain formats for export, especially NexML. (Currently only PNG can be exported. MathML files can be saved as well.)
- Ability to save just the definitions or just the phylogeny to a MathML file.
- Ability to import definitions from a MathML file.
- MathML tweaks. (Use csymbol instead of ci for taxa. Normalize presentation.)
- Ability to write in Newick strings directly.
- Skin various components (sliders, steppers, checkboxes, etc.).
- Fix line breaks in MathML formulas.
- Various scrollbar issues.
- Special character issues.
- Rich editor for taxon labels, including ability to edit taxon URIs.
- Arc bisection tool.
- Fix node merging (i.e., synonymization).
- Add ability to select definition type when creating a name.
- Node Pane Control Bar revisions. (Change Resolution Slider to a stepper. Add Zoom Slider.)
- Definition Editor tweaks/fixes. (Some actions are blocked that should be possible. Textual Editor does not always update. Various layout issues.)
- About/Help Panel.
12 May 2010
Why HTML 5 Canvas Will Not Be Replacing Flash That Soon
Not only are the arrowheads missing (as they are on Safari on all platforms, not just the iPhone), but the labels have bizarrely been placed outside the canvas, flipped upside-down! The tool works fine on Firefox and Chrome. (Internet Explorer has not implemented <canvas> yet, and I haven't played enough with the interim solution, ExplorerCanvas, to get it working.)
I think the <canvas> element is a cool idea, and I'll continue to play with it. But it has a long way to go to compete with a cross-platform tool like Flash. HTML 5 may be "open"—but it also needs to "work".
06 May 2010
PhyloPainter: Happy Little Trees
I'll probably write more on the topic later, but suffice to say for now that working HTML 5 feels like I've traveled in time back to 2001, the days of ActionScript 1.0. JavaScript is a poor language for anything complicated. Canvas has covered the basics of vector drawing well, but little else. That said, I see potential and I'm pretty certain the tools will improve.
For my first HTML 5 app, I ported some basic functionality from Names on Nodes, namely, the ability to read Newick tree strings and the ability to draw graphs. I give you:
It's a bit rough right now. For one thing, it doesn't work in Internet Explorer (despite the inclusion of a workaround JavaScript tool—the current version of IE doesn't support HTML 5 Canvas). But it's a start.
Give it a try—paint some happy little trees!